A non-technical summary of our project can be accessed here.
—
Specialised ribosomes are groups of ribosomes that specifically translate a set of mRNAs and represent a new regulatory mechanism in gene expression. This concept is relatively new, and the examples currently described are in single organisms and have limited mechanistic detail. Using cutting-edge scientific techniques, we aim to understand the detailed molecular mechanisms of specialisation and determine common features across eukaryotes.
We have selected examples of specialisation from our preliminary results and the literature, and will unearth additional novel candidates of ribosome specialisation through evolutionary analyses. All our results, and those from the literature, will be collated here.
Overall, this programme of research will allow us to unravel precisely how ribosome heterogeneity results in specialisation, and to define a ‘ribosome code’ applicable to all eukaryotes. In addition to advancing our knowledge of gene expression, our findings have the potential to impact our understanding of several human diseases. For example, ribosomopathies – including Diamond-Blackfan anaemia – are caused by mutations in ribosomal proteins, and it has been suggested that specialised ‘oncoribosomes’ might be involved in protein synthesis dysregulation in cancers. Additionally, understanding mechanisms of ribosome specialisation will enable the modulation ribosome translation in future medical, agricultural and biotechnological applications.
RiboCode will:
- Provide an unprecedented understanding of how specialised ribosomes regulate translation at the molecular level through changes in their composition.
- Reveal the critical sites of ribosome specialisation across eukaryotes.
- Generate a ‘ribosome code’ to explain the observed common mechanistic features of how specialised ribosomes regulate translation across eukaryotes.
- Employ the ‘ribosome code’ to predict additional routes of specialisation to provide the potential for future engineering of gene expression.
Themes
Our work is divided into four Themes; each with a key objective. Members of the RiboCode team will work collaboratively on each Theme.
Theme 1 Evolutionary genomics
Objective: Identify potential sites of ribosome specialisation and determine common mechanisms across eukaryotes.
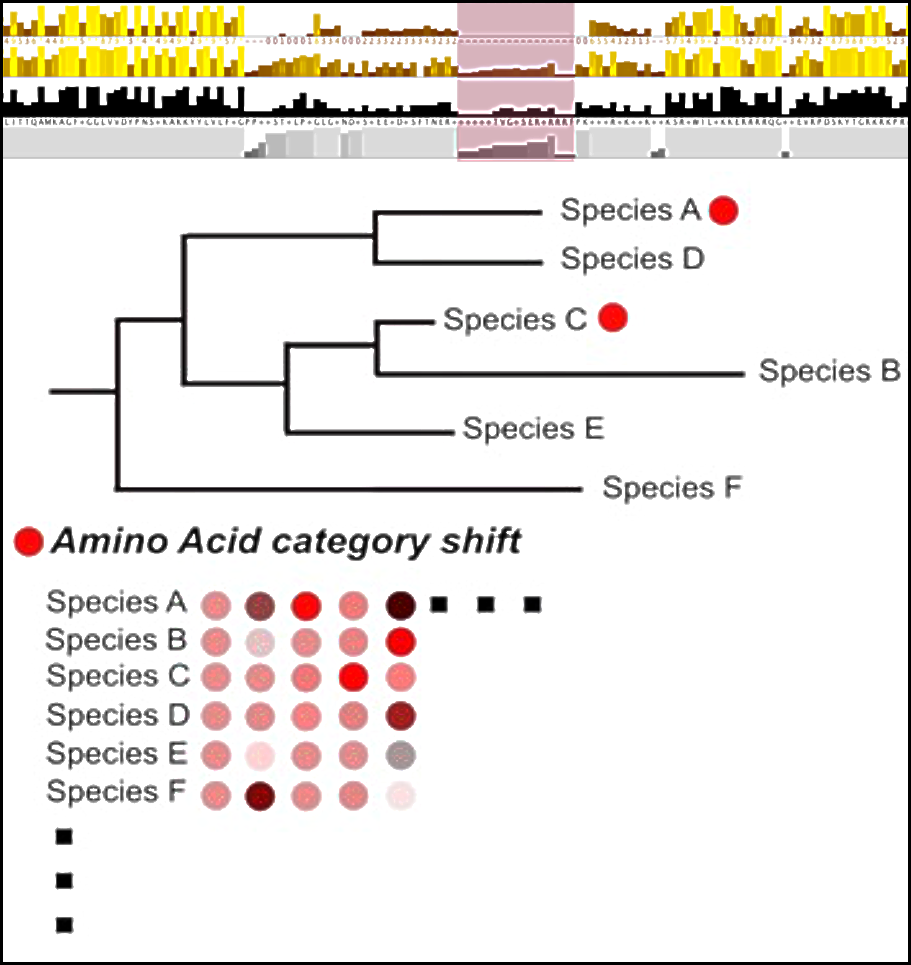
Theme 1 focuses on uncovering common routes to specialisation across all of eukaryotic life: cracking the “ribosome code”.
We will generate hypotheses of sequence-driven specialisation using conserved evolutionary features in RPs and associated proteins. This will be achieved using novel phylogenomics approaches designed for this project and large scale genome data analytics. We will extract key evolutionary features from the sequences of RPs and associated proteins identifying levels of conservation of these features across all evolutionary depths from eukaryote-conserved to Kingdom-conserved (plants, fungi, animals) and within Kingdom-conserved. Integrating data from across the program of research, and combining traditional comparative genomics and phylogenomics with machine learning techniques, we will generate hypotheses of ribosomal specialisation which the rest of the RiboCode team will test from a structural and functional perspective.
Theme 2 mRNA translation
Objective: Determine which mRNAs are translated by various specialised ribosomes.
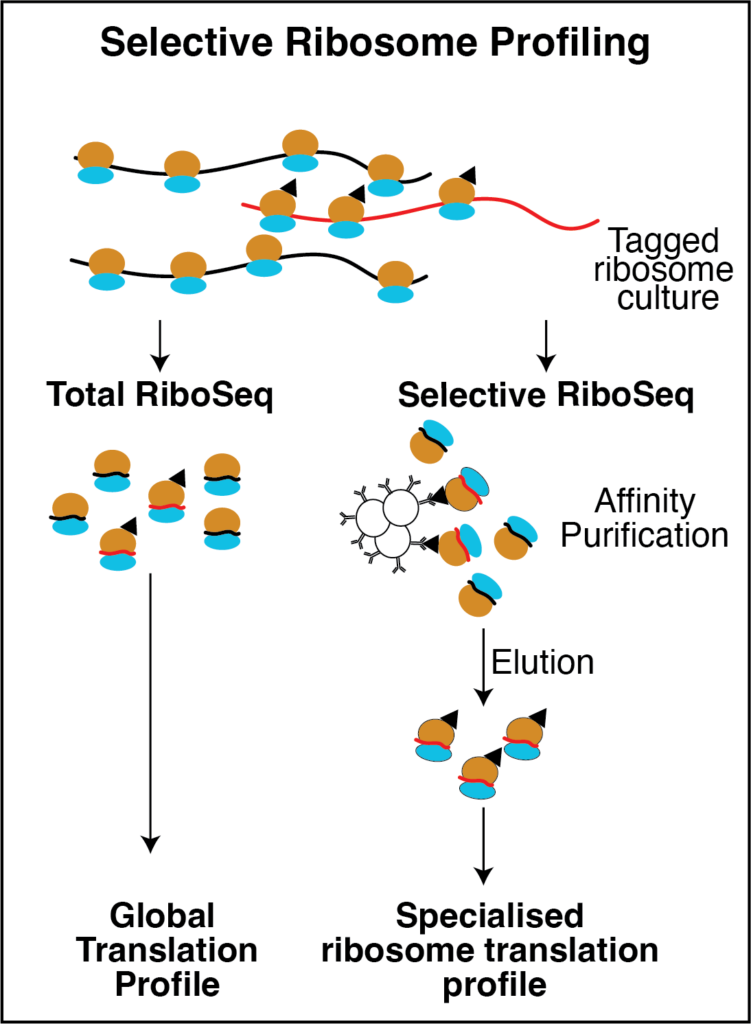
Theme 2 will investigate the functional and mechanistic conservation of specialised ribosome components in each of our five model systems. Central to both assessing a ribosome population for specialisation and dissecting their molecular mechanism of regulation is understanding specific translational output. To do this we will engineer cell lines and organisms with affinity tagged specialized ribosome components to enable selective RiboSeq. Riboseq will determine which mRNAs are regulated by specialised ribosomes and provide information on how this regulation may be achieved. This will allow the same “type” of specialized ribosomes to be compared across multiple organisms and provide experimental evidence of functional conservation. These analyses will also allow the identification of conserved features of mRNAs that are targeted by specialised ribosomes in each model, and inform subsequent functional testing. The role each specialised component plays in translation and its conservation will be assessed through mutational analyses, translational reporters and cross species complementation assays.
Theme 3 Structure of specialised ribosomes
Objective: Assess how variations in ribosome composition structurally regulate translation.
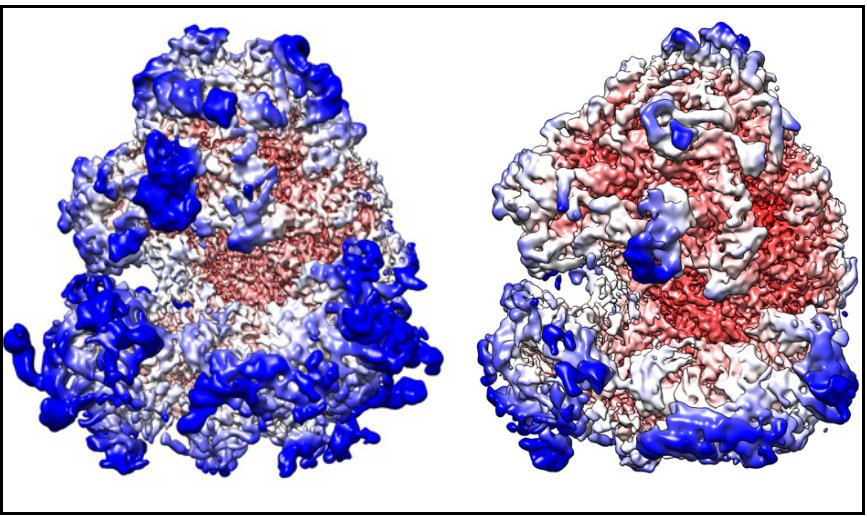
Central to dissecting the common mechanisms of translational regulation by specialised ribosomes, and something currently missing for most specialised ribosomes, is an understanding of the structural impact of specialisation on translation regulation. This will be addressed in Theme 3, which, when combined with Themes 2 and 3, will allow us to unravel the molecular mechanisms of ribosome specialisation.
Our established structural approach to characterising specialised ribosomes involves: 1) purifying candidate ribosomes and determining their protein composition, including the identity and stoichiometry of protein subunits that confer specialisation, by quantitative MS (e.g. TMT-MS), and 2) characterising their structure by cryo-EM, to provide insight into the structural differences that drive specialisation. This workflow has allowed us to unravel composition and structural differences in ribosomal paralogs between Drosophila ribosomes isolated from testes and ovaries, and we will use it during the sLoLa to gain structural information on the specialised ribosomes we will purify. To obtain further information on regions of the ribosome that have been historically challenging, we will adapt structural MS approaches for the study of ribosomes, including native MS, which allows unravelling the stoichiometry and composition of specialised ribosomes, and cross-linking MS, which allows probing for structural/dynamical changes in the ribosome.
Theme 4 Transformative technologies
Objective: Develop novel biophysical tools to probe ribosome specialisation.

Theme 4 will focus on developing a range of new biophysical tools to probe ribosome specialisation. For example, we will develop novel tools to enhance the functional analysis of ribosomal complexes and to complement and enrich the structural characterisation delivered by Theme 3.
One of the challenges in dissecting specialised ribosomes is small sample size, given their tissue/developmental time-point specific formation. We will develop a solid-state nanopore platform to tackle this limitation by enabling ribosome analysis with single entity resolution (Raveendran et al, ACS Sensors, 2020).
To test the ability of specialised ribosomes from one species to regulate translation in another, we will develop a nanoinjection platform based on nanopipettes to introduce ribosomes into naïve cells to investigate any translation reprogramming using state-of-the-art single cell proteomics approaches.
Finally, we will generate a database platform which integrate the data types for specialised ribosomes in different species across all WPs. The database will be made freely available to the scientific community to maximise the impact of our results.
